Photo : Purabi Deshpadne / Research Matters
Biology, a vast discipline, is today no longer restricted to studying living organisms in the wild, or working with cells and protein solutions in the confines of a lab. Interdisciplinary influences from mathematics, physics, chemistry, and statistics on biology are increasing by the day in relentless attempts to comprehend the complexity that is ‘life’. Bioinformatics is a result of one such effort, where computational approaches are used to solve problems that are practically either impossible to resolve or are highly time-consuming to pursue. Bioinformatics provides analytical solutions in the form of computer programs that trained researchers can use to interpret data.
With an increasing understanding of the subject and use of interdisciplinary approaches, are we now better equipped to fight near-fatal diseases like cancer? Every day, numerous research labs are generating massive bodies of knowledge in the form of research data and publications that has helped us understand the different tenets of this disease like never before. How, then, can we scoop out specific bits of research evidence or nuggets of information from the enormous number of research papers that can help us tackle cancer better? Using bioinformatics is what Ms. Sunita Yadavalli and Dr. Prashant Kumar from the Institute of Bioinformatics in Bangalore, propose.
“It is imperative to note the importance of data resources available online,” says Ms. Yadavalli. “It is possible to infer and deliver important insights from exhaustive data curation and without using bench resources. Particularly in the realm of cancer, any information is valuable information as it has not been elucidated completely”, she adds.
In a recent study, Ms. Yadavalli and Dr. Kumar, along with a team of researchers from the Indian Institute of Science and the National University of Singapore, have attempted to figure out what molecules and mechanisms are involved in the movement of cancer cells. Here is the interesting aspect of the study -- the researchers have analyzed cancer cells and the genes involved in the process using just a computer!
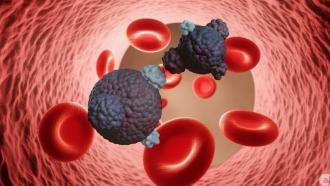
Cancer starts as tumors – a solid clump of cells that are static. Circulating Tumour Cells (CTCs) are cells that break free from the tumors and squeeze their way through the walls of the nearest blood vessel and enter the bloodstream. While most cancer cells don’t have the inherent ability to migrate, they acquire this ability through a cellular mechanism called Epithelial Mesenchymal Transition or EMT. This mechanism kicks in when certain genes in our cells are turned ‘on’. In the case of CTCs, the tumor cells turn on specific genes that allow them to change from being sedentary to being nomadic.
CTCs play a great role in cancer diagnosis and prognosis since they can be collected by simply drawing some blood from the patient as ‘liquid biopsy’. However, isolating and detecting CTCs in blood is not as easy as it may sound and is limited to a handful of validated techniques. This is because CTCs are not yet completely characterized at the molecular level– we do not know enough about them to easily differentiate and pick these cells from all the other normal cells in the blood.
Ms. Yadavalli’s work involves examining the cellular pathways involved in the migration of tumor cells into the bloodstream, a study that could help in cancer diagnosis and therapy. During the study, she scoured through thousands of research papers by scientists who had already worked in the lab and had generated large sets of data using different experiments. She then narrowed this down to five studies where the scientists used one or more advanced techniques in their isolation of CTCs to overcome the problem of contamination. The diverse data set that she began with had data from CTCs obtained from skin, prostate, breast, colorectal and pancreatic cancer patients.
After a thorough analysis, the researchers found that to move into a blood vessel, CTCs tend to switch on the same genes that white blood cells (a.k.a. leukocytes) normally use to enter and exit the bloodstream. “However, when we investigated further, we saw that across cancers, these cells turn on various genes that have already been categorized under the leukocyte extravasation pathway and more. After that, everything fell into place, and we could weave an insightful research article”, comments Ms. Yadavalli.
The study, a first of its kind that used just data from previous CTC research, reveals a number of key molecules that can be exploited for both diagnosis and therapeutic purposes for cancer. Although there has been research in the past that attempted to bioinformatically characterize EMT in cancer cells, these studies involved cells from patient-derived tumours or ones that were growing in the lab; this is the first study that characterizes the pivotal EMT in CTCs isolated from patients. It needed no expensive machinery or pricey chemicals, or for that matter, not even a lab!
The power of analyzing data, thanks to bioinformatics tools, provides valuable interpretations that vary from study to study, thus helping us gain new perspectives and insights that the researcher who generated the data wouldn’t have realized. “With big data that exists today in the scientific research community, all you need is a computer, basic bioinformatics know-how and the dedication and inclination to process large data sets. You can study complex biological phenomena and come up with answers that are beyond the scale of what a wet-lab can provide”, signs off Ms. Yadavalli.