Scientists at Centre for DNA Fingerprinting and Diagnostics (CDFD) here studying salivary microbiome diversity feel the findings can help in early diagnosis of certain diseases by understanding the perturbations to this microbiome composition.
Dr. Madhusudhan R. Nandineni, of CDFD and scientists from the Max Planck Institute for Evolutionary Anthropology, Germany have been studying the microbes present in the saliva of Indians. Funded by Department of Science and Technology (DST), Government of India and the Max Planck Society, the findings of the study were published in the journal PLoS ONE.
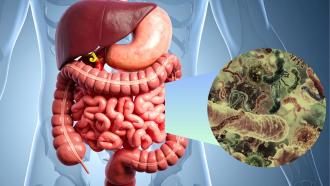
Microbes in our body help in the digestion of food, production of vitamins and enhancing immunity in the host. Studies have shown that an imbalance in the composition of these microbes could result in liver diseases, gastrointestinal cancers, oral disease, psychological diseases and other autoimmune diseases.
“The microbiome or the small environment in which microbes thrive, of an individual comprise both friendly and detrimental bacteria. The study provides a snapshot of salivary microbiome diversity among healthy individuals and aids in deciphering the role of the onset of certain diseases by understanding the perturbations to this microbiome composition”, says Dr. Madhusudhan.
To decipher the microbiome, the researchers studied a total of 92 saliva samples of healthy volunteers from 8 different states in India spanning North India (Jammu & Kashmir and Uttarakhand, East India (Jharkhand, West Bengal and Assam) and South India (Andhra Pradesh, Telangana and Tamil Nadu).
Once the samples were collected, the researchers made thousands of copies of a segment from the 16S rRNA gene -- a widely used gene to establish taxonomic relationships between different bacteria. These gene sequences were analyzed in two ways -- one by directly identifying the bacteria using public databases, and the other by grouping the sequences based on their similarities into 'operational taxonomic units' (OTUs). The researchers identified 785 such OTUs that included 165 bacterial genera in Indian populations.
The study found that Indian population had more and unique set of bacteria than their African, Alaskan and German peers. Within Indians, the North Indian population had more variation in the bacterial genera, while South and East Indian populations come second and third. However, the diversity of bacteria genera within individuals of same regions was comparatively less.
The researchers identified a wide variety of bacterial genera including Streptococcus, Prevotella, Fusobacterium, Veillonella, Leptotrichia, among others. Among these, the researchers found that Streptococcus was predominant with the North Indian sample at 35%, the East Indian sample at 42% and the South Indian sample at 38%. Samples collected from Tamil Nadu commonly had Atopobium, Megasphaera and Prevotella, whereas Streptobacillus and Bacillus were common in the Telangana samples. Also, Neisseria and Streptobacillus were significantly high in the South Indian samples.
Among 785 OTUs identified, all the three regional samples had 660 common OTUs, and all individuals had 37 OTUs in common. The study also found that geographical location was the major factor that contributed to the diversity in the microbes present in the saliva. The findings correspond with previous studies that point out to similar food habits between populations, leading to lesser diversity in microbial composition.
The study also contributed 9 new bacterial genera such as Meiothermus, Hydrocarboniphaga, Streptobacillus, Brevibacterium, Brevibacillus, Weissella, Aeromonas,Chromobacterium and Aeribacillus to the 'Human Oral Microbiome Database' (HOMD). “This work emphasizes the importance of studying underrepresented populations like India and aids in understanding their occurrence with relation to India's geography, food habits and lifestyle of the contemporary populations”, says Dr. Nandineni.
Indeed 'unity in diversity' describes India not only for its people, but also about their salivary microbiome!