Researchers identify molecular markers that can help in developing better varieties of black pepper
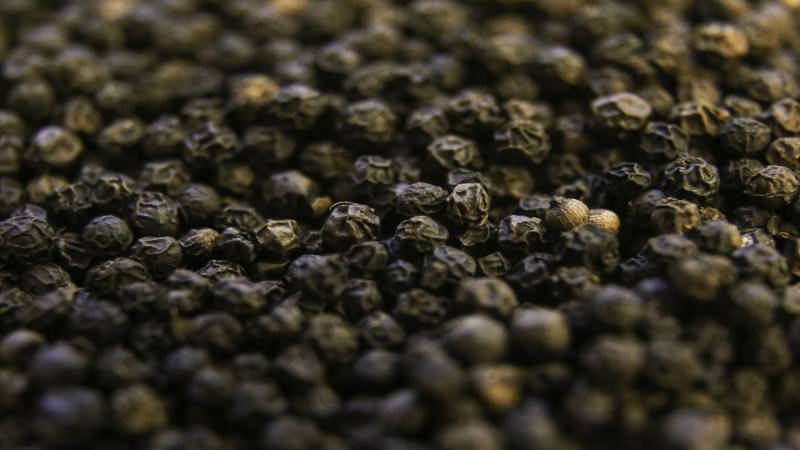
Dubbed as the 'king of spices', black pepper has a lot to offer—whether as a flavouring agent or as a traditional medicine to relieve common cold, chest congestion or sore throat. A vital ingredient in many cuisines, it is the only spice that finds its way on nearly everything on the dining table. Piperine, a chemical in the black pepper, gives it a sharp taste. It is a native plant of the Western Ghats, and the crop is of immense agronomic value in today's world trade. However, insufficient genomic resources pose as main barriers in cultivating disease-resistant varieties of this master spice.
In a recent study, researchers from ICAR-National Bureau of Plant Genetic Resources, New Delhi, have identified a type of DNA sequence, called Simple Sequence Repeats (SSRs), which aims to ease and promote the genetic analysis of black pepper. The findings of the study, published in the journal PLoS ONE, could lead to enhanced productivity with better traits of the plant.
Simple Sequence Repeats, also called microsatellites, are repeating stretches of DNA that contain one to six nucleotide bases, and are randomly present throughout the genome. These can be passed from one generation to the next. Hence, they are used as a 'signpost' to keep track of a gene of interest. They play critical roles in genetic studies and plant breeding. The location of these SSRs in the genome remain the same in related species, thus enabling cost-effectiveness of similar studies on them.
Unlike other commercial crops like watermelon, cotton or finger millets, very few genetic studies have been done on black pepper, due to lack of resources. Although the Western Ghats harbours the maximum genetic diversity of black pepper, it remained "largely untouched from genomic interventions," say the researchers. The identification of cross-species SSRs will save time, effort and resources in the development of SSRs in these species, and aid future genetic and evolutionary studies.
The researchers first sequenced the entire genome isolated from the leaf samples of black pepper (Piper nigrum) and then scanned it for the presence of SSRs. The genome was then amplified with a method called Polymerase Chain Reaction, using short nucleic acid sequences called primers, designed to identify the right SSRs. The researchers compared the results of their analysis in thirty different types of black pepper. The researchers also explored ten species of Piper (including P. nigrum) for the presence of these SSRs by looking at an online database of the genetic sequence.
In all, 69,126 SSRs were identified in the study, with the majority of SSRs composed of two nucleotide repeats—thymine and adenosine (TA). Out of 85 primers, 74 produced the required results of appropriate-sized SSRs. Genetic diversity study of the thirty cultivars reported four distinct groups. A few SSRs were found in closely-related species, implying that they could be used in studying other species of black pepper with limited genomic knowledge.
The current study seeks to fill the void in genomic knowledge of black pepper species by identifying easy-to-detect molecular markers to enable its genetic and breeding studies. The SSRs, which indicate the location of a gene related to the desired trait, can help in choosing the right plants for breeding experiments, which is otherwise a difficult activity.
"The genomic microsatellite markers identified in black pepper in this study would form valuable and long-awaited resources for researchers and plant breeders", say the researchers.
The findings can also be used for diversity studies, linkage mapping, evolutionally biology, DNA fingerprinting and trait association shortly.