Researchers use a data-driven approach to identify bat species that could be carriers of the Nipah virus in Kerala.
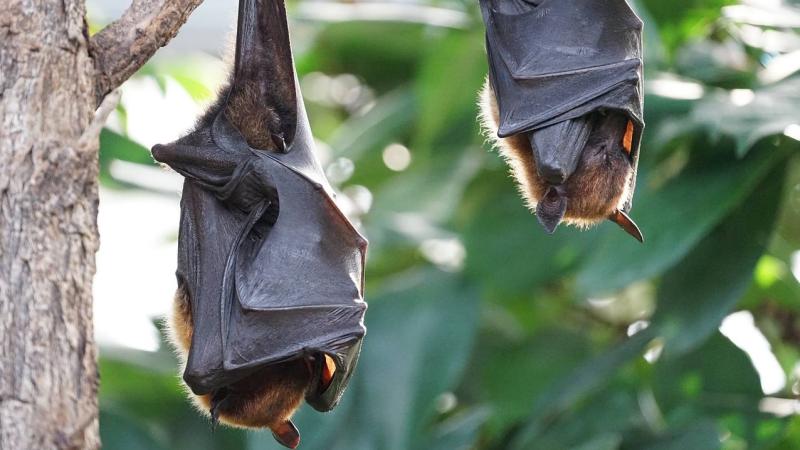
On May 2, 2018, Kerala recorded its first case of the Nipah outbreak—a near-fatal viral disease caused by the Nipah virus. Characterised by symptoms that include fever, headache, drowsiness and mental confusion, the illness claimed 16 lives. Bats, known to be the carriers of the virus, had passed the virus to humans. Although studies initially pointed at insectivorous bats as the culprit, the virus was later found in Indian flying fox, Pteropus medius, which feeds on fruits and nectar.
The outbreak of Nipah in Kerala was not the first in India; isolated cases have been reported from West Bengal in 2001 and 2007, attributing the source of the infection to Bangladesh, where it recurs year after year. “Drinking date palm sap or eating fruit contaminated by bat urine or saliva increases the risk of becoming infected by Nipah virus”, says Dr Barbara Han, Disease Ecologist at the Cary Institute of Ecosystem Studies, in an interview with Research Matters. However, are there some species of bats that are more likely to carry the virus than others?
In a new study, Dr Han and other researchers from institutes in the USA and India, have tried to use machine learning to answer this question. By analysing previous scientific records of Nipah virus and its prevalence around the world, they have identified eleven species of insect- or fruit-eating bats that pose a high risk of transmitting Nipah virus in India Their findings are published in the journal PLOS Neglected Tropical Diseases.
The Nipah virus belongs to a class of viruses called henipaviruses, found in bats throughout Asia, Africa, Australia and the Americas. Very few studies detail what viruses of this class are found in which bats and where, and the patchy data poses a challenge to get an overall picture on the prevalence of the infection.
The researchers of this study searched through repositories like the Web of Science and PubMed for published studies on henipaviruses in wild bats. Out of the 285 records and 25 papers found on Nipah, they recorded the species, the country of sampling, the sample size and the technique used to identify the presence of the virus. They used data about some of the traits of these confirmed carrier bats to predict other species that could be carriers, but not yet identified thus.
“Traits include features that are intrinsically different between species, like their body size, diet and the size of their geographic range”, explains Dr Han.
The researchers trained a machine learning algorithm with data of 48 traits of 523 bats found in Asia, Australia and Oceania. Their algorithm 'learnt' the associations between the characteristics of these bats and whether they tested positive for the virus. Out of the 112 bat species that occur in India, the algorithm identified 11 species as confirmed carriers of Nipah virus, which had evidence of past or current infection in their blood, urine, or tissue.
The study identified seven bat species from Kerala as 'reservoirs' of Nipah virus. These included the Lesser short-nosed fruit bat (Cynopterus brachyotis), Greater short-nosed fruit bat (Cynopterus sphinx), Cave nectar bat (Eonycteris spelaea), Leschenault's rousette (Rousettus leschenaultii), Indian flying fox (Pteropus medius), Lesser Asiatic yellow bat (Scotophilus kuhlii) and Pomona roundleaf bat (Hipposideros pomona). Out of these, Pteropus medius is the only species that had Nipah virus isolated from it, while the rest showed antibodies against Nipah in their blood.
“Pteropus medius is fruit/nectar eating bat that shares many features in common with other bat species and also coexists with several other species in regions where Nipah virus spills over regularly”, says Dr Han.
Besides, the study also identified six species of bats whose geographic ranges overlapped with Asia, Australia and Oceania. These bats were likely to carry the virus, but are not yet identified as Nipah reservoirs. Since many of the traits of these bats resemble that of the known carriers, the researchers believe they have a high likelihood of being carriers of the virus. Species like the Long-winged tomb bat (Taphozous longimanus), and the Black-bearded tomb bat (Taphozous melanopogon), found in Kerala, may have an 80% probability of exposure to the virus, they say.
The study identifies a list of species that need to be on our 'watchlist' to address future outbreaks of Nipah, which does not yet have a vaccine. With the World Health Organisation (WHO) identifying Nipah as a likely epidemic, controlling the spread of the virus is vital to minimise the devastating loss of lives.
"The first step to preventing spillover transmission of any animal-borne infection is to identify which animals may be serving as sources (reservoirs) of infection. Our study applies a data-driven approach to identifying those animals", says Dr Han, talking about the implications of the findings.
However, this list of species is not exhaustive, warn the researchers, as global phenomena like climate change and global warming can impact the availability of food for bats and their immune system, making more species susceptible. "When bats don't have high-quality food sources, or when their natural food sources are depleted due to urbanisation, they switch to sub-optimal food that is more reliably available and become nutritionally stressed. This can lead to higher amounts of the virus being shed", explains Dr Han, highlighting the increased risk of us contracting the disease.
As the next step, the researchers plan to check for the prevalence of the virus in more species and match the data with predictions of their algorithm. "We hope these results will contribute to better understanding where and when the risk of spillover to humans is greatest, and how that risk may change over time," concludes Dr Han.
This article has been run past the researchers, whose work is covered, to ensure accuracy.