If you have been to the movies or paid a little attention to the advertisements shown on the television, you may be familiar with the infomercial depicting the ill effects of smoking and chewing tobacco. Whenever a character smokes, the message that “smoking causes cancer” flashes on the screen. However, despite these warnings and the gruesome pictures on the cigarette packs, about 266.8 million Indians still used tobacco in 2018.
Time and again, studies have linked smoking and chewing tobacco to oral cancer and other conditions leading to death. However, the molecular mechanisms that occur in our cells and result in cancer have not yet been completely deciphered. In a recent study, researchers at the Indian Statistical Institute, Kolkata; University of Otago, New Zealand; Fred Hutchinson Cancer Research Centre, USA and National Institute of Biomedical Genomics, West Bengal, have explained how altered levels of biological molecules, like lncRNA and miRNA, can lead to the development of oral cancer.
MicroRNAs or miRNAs are small RNA molecules that target messenger RNA molecules and might prevent their conversion into proteins. Though vital to the cell for protein synthesis and gene regulation, miRNAs have been associated with diseases like cancer. Their levels in the cell sometimes fluctuate due to various factors, including DNA methylation, where a methyl group (-CH3) is added to the DNA. This addition modifies the DNA, thereby affecting gene expression and function.
In the current study, published in the journal Epigenomics, the researchers have explored how methylation can cause a difference in the levels of miRNA. Methylated DNA strands consist of both methylated and unmethylated cytosine molecule, one of the four main base molecules found in the DNA along with adenine, guanine and thymine—commonly referred to as A, G, C, T, respectively.
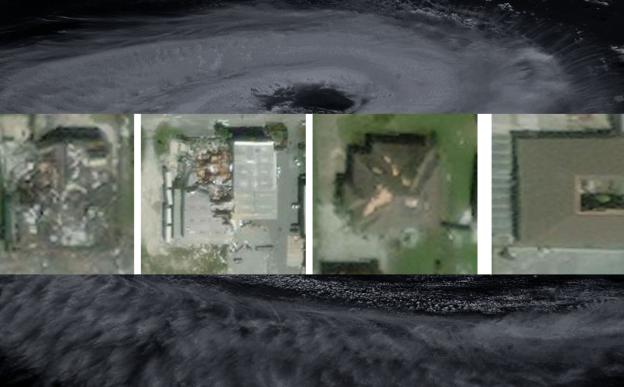
Scientists study DNA methylation using a technique called bisulfite conversion, which converts all unmethylated cytosine molecules into uracil, while leaving methylated ones intact. The researchers of the current study used this technique to analyze the methylation patterns in the genome from tumour samples obtained from the Guru Nanak Institute of Dental Science and Research in West Bengal, in collaboration with Prof. Ranjan. R. Paul and Prof. Mousumi Pal. They then compared the tumor samples against healthy tissue samples and generated lots of data about these patterns.
“For this study, we had huge DNA sequences to analyze; the size of which was a challenge and the task was expensive and laborious. Hence, we restricted our sample size to 16, which included the analysis of 16 tumours and their paired normal tissue samples, obtained from 16 patients”, explains Prof. Bidyut Roy from the Indian Statistical Institute, Kolkata, who was the lead author of the study.
The researchers found that in the tumour samples, many hypermethylation areas were closer to sites in the DNA where the synthesis of miRNA began. Short stretches on DNA, called CpG islands, where there is a high frequency of cytosine and guanine molecules, are the ones that usually undergoes methylation. The study found that the cores of the CpG islands were highly methylated, whereas the sites on their either sides had low methylation. Generally, hypermethylation at miRNA genes leads to reduced levels of the miRNAs.
When the gene sequencing data was narrowed down to 144 unique miRNAs, more than 80% of them showed high levels of methylation. About 46 of them were intergenic miRNAs. Tumour growth and migration, two of the characteristics of cancer cells, were found to be activated by these 46 miRNAs.
The study also proposed that hypermethylation of specific miRNAs was also associated with poor survival of the patients, showing miRNAs indeed play important roles in regulating genes responsible for the suppression of cancer. The researchers also analysed the methylation patterns in cancerous tissues from smokers, tobacco chewers and mixed habits; and found that 21 unique miRNAs were differently methylated in smokers, 8 in tobacco chewers and 76 in those who smoked and chewed tobacco.
The researchers believe that oral cancer associated with tobacco chewing habits should be the focus of further studies since this habit is predominant in south-east Asian countries. As a next step, they are now focusing on studying the methylation patterns of genes responsible for protein synthesis.
This article has been run past the researchers, whose work is covered, to ensure accuracy.