The Golden age of antibiotics was an era spanning from 1950 to 1960, during which most common antibiotics like Penicillin, Streptomycin, and Chloramphenicol were discovered. These antibiotics were considered to be 'silver bullets' that would soon eliminate all bacterial infections. Unfortunately, bacteria proved to be extremely resilient, and resistance to these drugs quickly emerged. Today, bacterial pathogens resistant to nearly all antibiotics threaten to undo decades of medical progress. The indiscriminate use of antibiotics for the treatment of common infections and in the food industry have exacerbated the problem, giving rise to new drug-resistant pathogens worldwide. Hence, researchers from around the world are compelled to discover better antibiotics to kill some of the most potent drug-resistant bacteria. In one such attempt, researchers from the Indian Institute of Science, Bengaluru, led by Prof. Nagasuma Chandra, have used a computational approach to design and synthesise amino acid polymers called peptides, which are highly effective in killing an array of multi-drug resistant bacteria.
Hundreds of such peptides, commonly called as 'antimicrobial peptides' (AMPs), that fight against invading bacteria, are found in multicellular organisms, including humans. The researchers of this study, published in Journal of Biological Chemistry, used a machine learning algorithm that studied the sequences of thousands of such peptides. They deciphered the common patterns unique to those peptides. As an analogy, a meaningfully constructed sentence inherently uses a framework of rules that can be considered as its grammar. The researchers looked for such a 'grammar' in the sequences of these antimicrobial peptides.
“Protein sequences remarkably are similar in structure to language, where complex rules about syntax and grammar help define the semantics (or meaning) of a sentence", says Deepesh Nagarajan, from the Department of Biochemistry, IISc, the first author of the study. To reveal this hidden grammar, the researchers used a neural network-based algorithm called Long short-term memory (LSTM). “LSTMs have a kind of ‘memory’ that allows it to retain information for 1000s of time-steps, which is very useful for tasks such as speech-recognition and language processing”, remarks Tushar Nagarajan who wrote the algorithm.
The researchers trained their algorithm by feeding it with thousands of unique peptide sequences that were known to possess antimicrobial properties. The training helped the algorithm to analyse and learn. “We had thousands of peptide sequences to analyse, whereas, at the time of previous work, only hundreds of antimicrobial peptides were characterised”, says Deepesh, commenting on the success of the prediction. The program then predicted several thousand candidates that had potential antimicrobial activity.
A few antimicrobial peptides were synthesized and found to possess activity against a variety of common pathogens. The best peptides were found to inhibit multi-drug resistant pathogens resistant to drugs of last resort like meticillin and carbapenem antibiotics. Further, one of the peptides tested was able to decrease the bacterial load of drug-resistant Acinetobacter baumannii a hundred-fold in 4.5 hours, in a mouse model of infection.
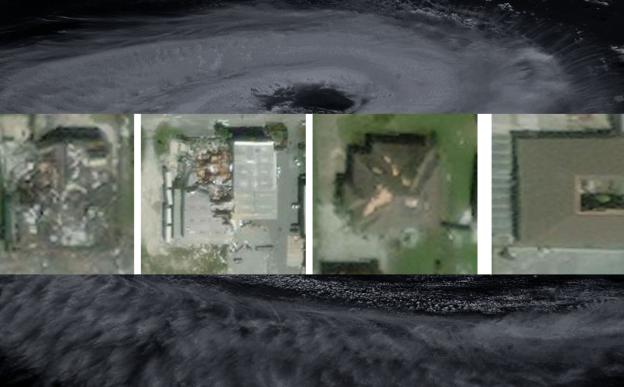
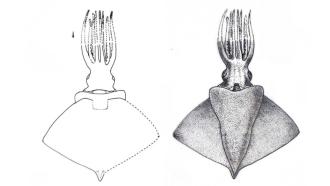
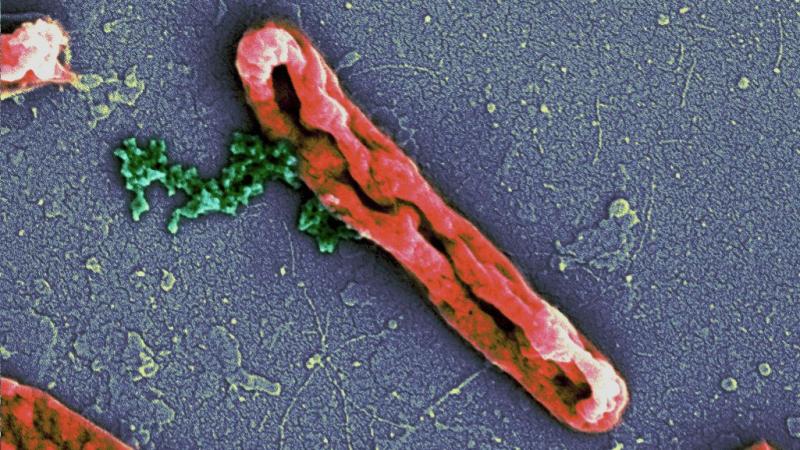
So how did the peptides act on bacteria? "We discovered that the peptides bound to and disrupted the bacterial cell membrane" says Prof. Disphikha Chakravortty, an author of the study. Peering through an electron microscope, they saw that the cylindrical bacteria had become ridged and flattened after being treated with the peptides. The bacteria were seen bleeding out cytoplasm through the holes created by the peptide binding. Since the entire cell membrane is targeted, the researchers believe that the bacteria will never be able to develop resistance against it.
“By targeting and disrupting the entire cell membrane, we simultaneously target a variety of lipid molecules, membrane proteins, and ion channels. The pathogen has to simultaneously evolve changes in all these molecules to develop resistance, which is much more difficult to accomplish via random mutations”, says Nagasuma Chandra.
This path-breaking study succeeds at developing drugs that target the bacterial cell as a whole, rather than focus on a single molecule within the cell. Since drug-resistant bacteria are quick to evolve, any bacterium in the community with a slightly different version of the targeted molecule has the potential to survive and thrive. The peptide molecule, developed by the researchers causes irreparable damage on a scale that the bacterium cannot recuperate, let alone develop resistance!