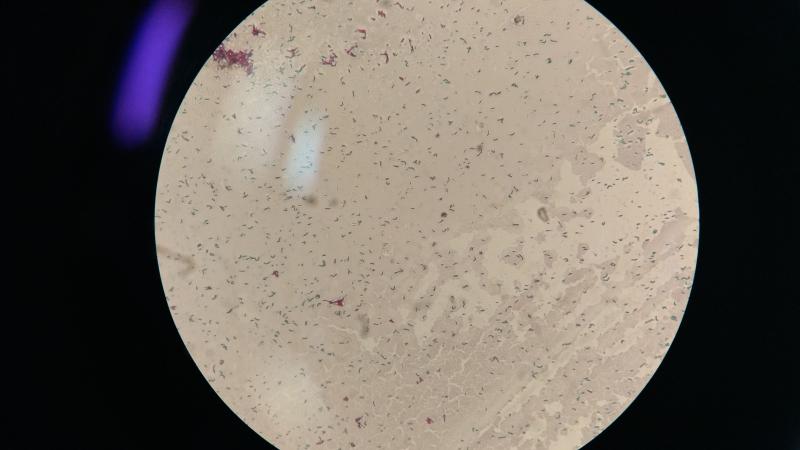
On the evening of March 24, 1882, German scientist Robert Koch discovered Mycobacterium tuberculosis--a tiny, rod shaped bacteria that divides after every 15-20 hours and causes a catastrophe called tuberculosis (TB). 135 years have passed since and scientists around the world are still struggling to find the perfect drug that can effectively combat this bacteria, which has consumed 10.4 million lives across the globe in 2016 alone. Though antibiotics help to an extent, the challenge that scientists face is that TB drugs fail to cure the disease effectively. Now, scientists from the Central University of Gujarat (CUG), Gandhinagar, are using computers to design the most efficient drug against TB.
The process of creating any novel drug involves screening a large number of synthetic and natural compounds and selecting suitable candidates or lead compounds, which kills the pathogen. As the number of such compounds grows each day, manually screening them is impossible. Hence, scientists are now turning to computational methods to help them solve this problem, and they call this ‘pharmacophore modelling’.
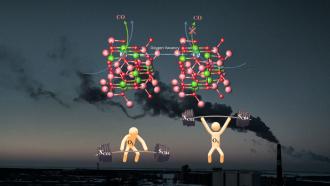
“It is a well established fact that the drug discovery and development is a time-consuming and costly procedure. Therefore, application of computer-aided molecular design and development of computational methods for lead generation and optimization are of enormous significance to reduce the overall cost and cycle time allied with drug discovery research”, says Dr. Prakash Jha from CUG. In a recent study published in the Journal of Theoretical Biology, Dr. Jha and his colleagues applied a pharmacophore modelling approach to understand the interactions between proteins found in Mycobacterium tuberculosis (Mtb) and the compounds which can suppress the bacteria.
The scientists first selected protein-ligand complexes from an online database. In this case,the protein belongs to the bacteria and the ligand is part of the compound or chemical molecule which, when binds to the protein, is capable of affecting the protein’s function. Then, they looked for similarities between these complexes. Using a software, they modelled accurate molecular structures of the protein-ligand complexes and screened them against a focused dataset in order to understand how efficient these models were. The researchers then performed enrichment studies to determine which molecules are active and capable of effectively binding to the bacterial proteins and destroying them. They then used statistical methods to group together similar models of molecules that could act against the bacteria.
The results showed that in order to make the ligands bind to the desired proteins, the researchers had to build more advanced models. It was found that the enzymes, purine nucleoside phosphorylase, 3-dehydroquinate dehydratase and ribose 5-phosphate isomerase were similar in their chemical features in three aspects--the closest observed in this study. These enzymes could work as drug targets in future, opine the researchers.
These hypotheses can be further employed in virtual screening.This will ultimately tell us what features of the ligand will help in performing efficiently against the proteins of Mtb. This study has also provided evidence that enzymes like ribose-5-phosphate isomerase are less explored as they are in the initial stage of development, and there is a dire need for advanced studies.
“This research provides a guiding principle for designing efficient inhibitor by taking into account the available pharmacophoric space. Moreover, this research is helpful for the upliftment of pharmacophore models that can be employed as an efficient pharmaceutical filter as well as for the design of a coherent inhibitor strategy that is effective against the bacteria”, says Dr. Jha about the importance of the study. While we have not yet found the wonder drug against TB, studies like this take us a step closer towards achieving it.